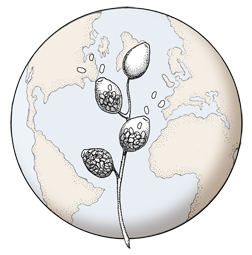
Scanu, B., Hunter, G. C., Linaldeddu, B. T., Franceschini, A., Maddau, L., Jung, T., Denman, S. (2013), A taxonomic re-evaluation reveals that Phytophthora cinnamomi and P. cinnamomi var. parvispora are separate species. Forest Pathology. doi: 10.1111/efp.12064
Between 2008 and 2011, severe dieback associated with root and collar rot was reported on Arbutus unedo in several sites in Sardinia, Italy. Isolations from infected tissues and rhizosphere soil samples consistently yielded a Phytophthora species. It was initially identified as Phytophthora cinnamomi var. parvispora Kröber and Marwitz by comparing morphological features with the original description and the internal transcribed spacer (ITS) sequences with those present in GenBank. A multigene phylogeny based on DNA sequence data from two nuclear (ITS and β-tubulin) and two mitochondrial (cox1 and cox2) gene regions combined with extensive morphological and physiological properties of these isolates, including the ex-type culture of P. cinnamomi var. parvispora, demonstrates that this taxon is unique and it is redesignated here as Phytophthora parvispora sp. nov. Although morphologically similar to P. cinnamomi, P. parvispora differs by its smaller-sized sporangia, chlamydospores, oogonia and oospores, higher oospore wall index, single-celled antheridia, higher minimum and maximum temperatures for growth and faster growth at optimum temperature. In the phylogeny, P. parvispora falls within Phytophthora ITS clade 7a, grouped in a well-supported clade sister to P. cinnamomi. In pathogenicity tests, P. parvispora and P. cinnamomi were equally aggressive towards A. unedo seedlings. The possible geographic origin of P. parvispora is also discussed.